
These, sometimes rather impressive, figures are made with the amazing YASARA
JRNL AUTH J.A.SPEIR,S.MUNSHI,G.WANG,T.S.BAKER,J.E.JOHNSON JRNL TITL STRUCTURES OF THE NATIVE AND SWOLLEN FORMS OF JRNL TITL 2 COWPEA CHLOROTIC MOTTLE VIRUS DETERMINED BY X-RAY JRNL TITL 3 CRYSTALLOGRAPHY AND CRYO-ELECTRON MICROSCOPY. JRNL REF STRUCTURE V. 3 63 1995 JRNL REFN ISSN 0969-2126 JRNL PMID 7743132 JRNL DOI 10.1016/S0969-2126(01)00135-6 |
To look at a whole virus and still be able to rotate it interactively, you need to use YASARA. And to calculate things on a virus you need WHAT IF.

|
These, sometimes rather impressive, figures are made with the amazing YASARA |
Lets first give some CCMV figures to illustrate what it looks like. In each case you can click on the figure to make it rotate. The first picture is just an all-atom model looked at from some 300 Ångstrøm distance.

|
Figure 1. The all atom model of CCMV. |
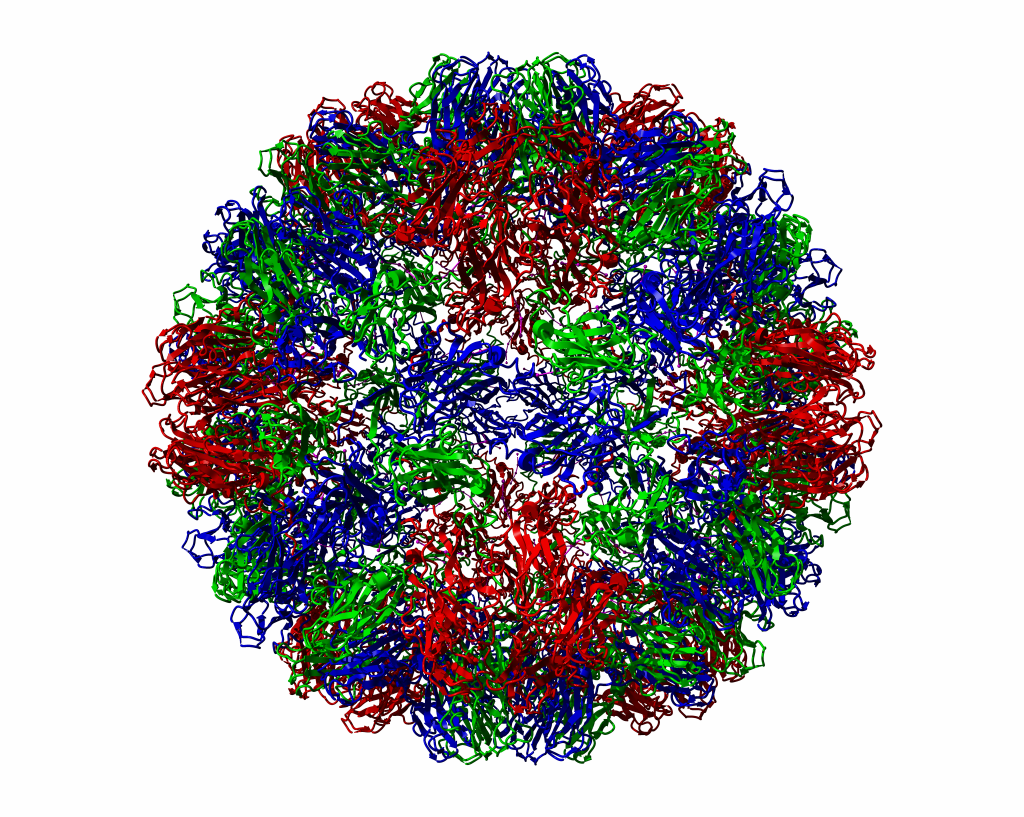
In the second plot we see CCMV as a ribbon-model, again from some 300 Ångstrøm distance.
|
Figure 2. |
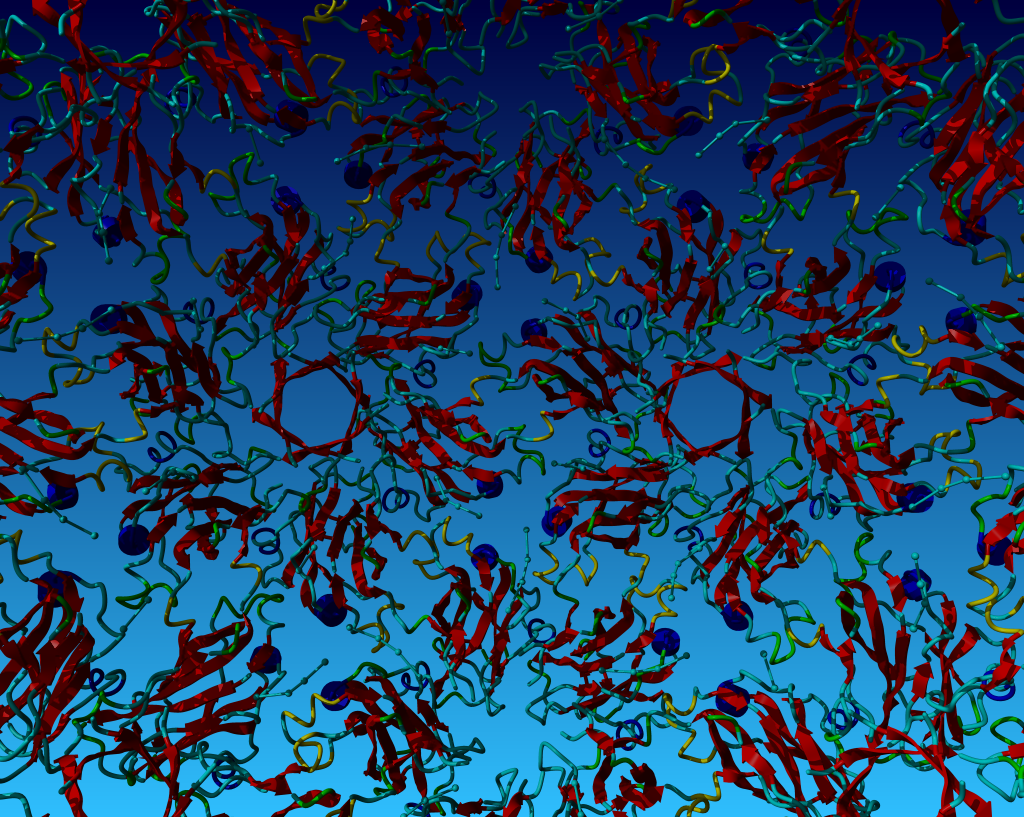
In the the next picture the camera was placed inside the virus and we see the shell from the inside rotate around us.
|
Figure 4. Strands in red, loops in blueish-yellow, helices in blue. The RNA is in purple. |
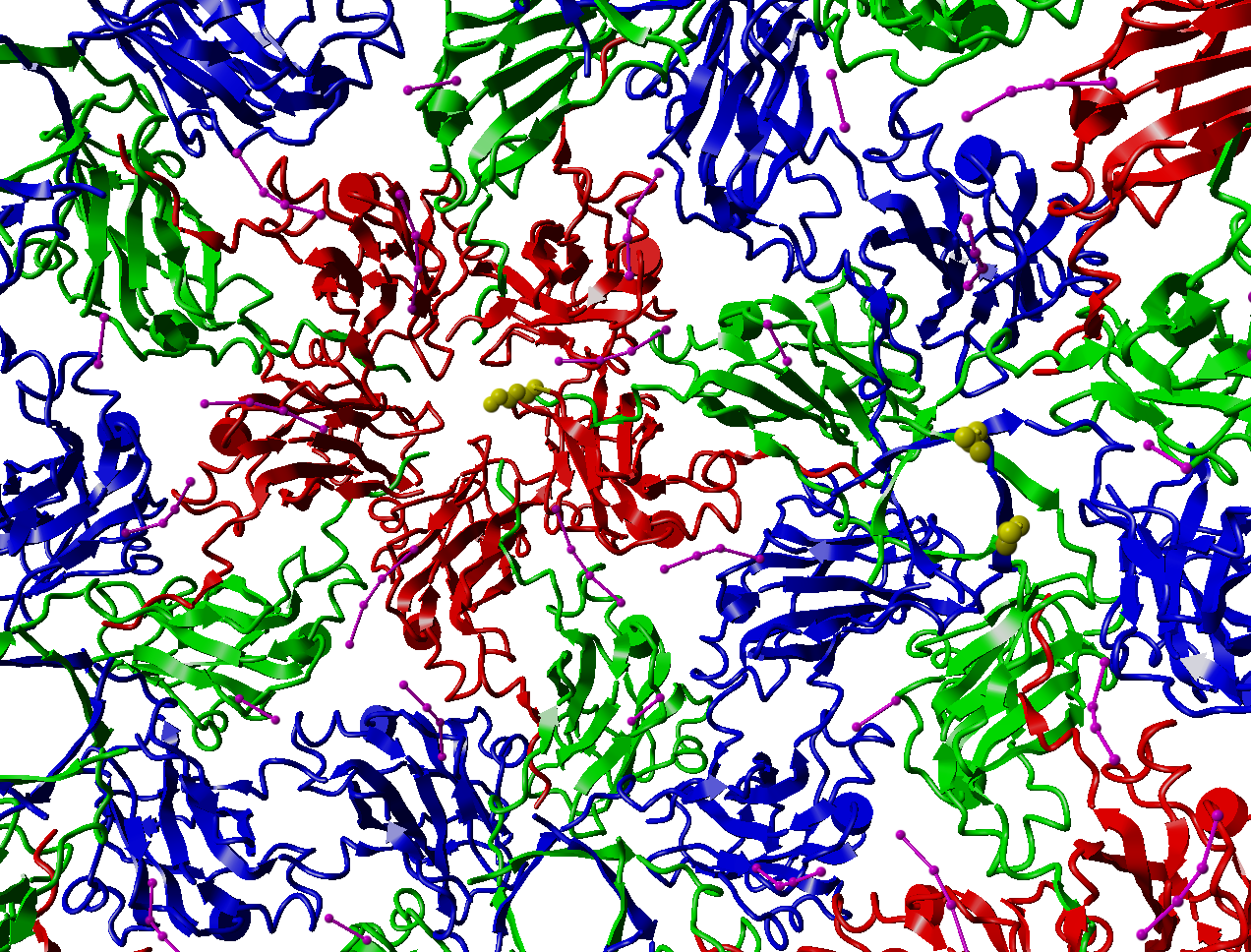
Something funny. If we look at the three C-termini, we see that the Ntermini of the green and blue units are very close to each other, but the N-termini of the red chains, that cluster around the 5-fold axis, are far away from the other two. When we were doing NMR on this virus in the early 80's, we never understood why the NMR signal for the N-termini was 'less than expected'. Now we know at least how to start thinking about this point.
|
Figure 5. Location of the N-termini of the three subunits indicated by showing the most N-terminal, X-ray visible, residue as a sphere-model. |