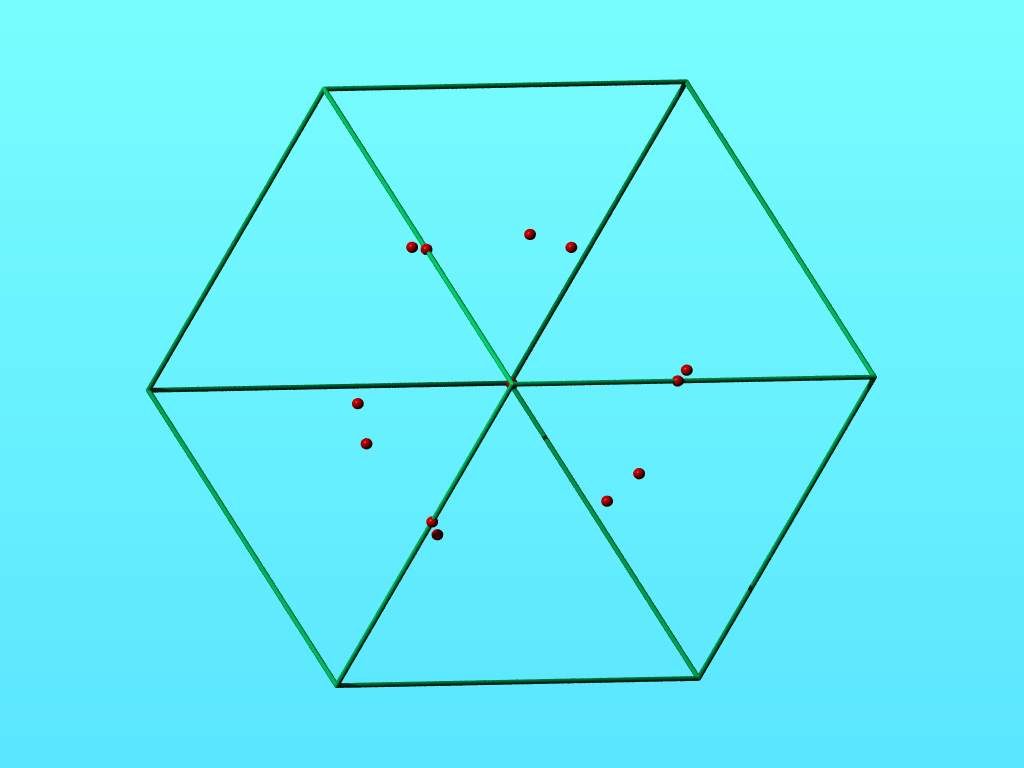
Waters with non-1.0 occupancy shown in a wireframe cell of 1QU9.
We all know that if an atom sits at a special position, i.e. exactly on a symmetry axis, then you should reduce its occupancy in the PDB file by the exact same factor as the multiplicity of that symmetry axis.
JRNL AUTH K.VOLZ JRNL TITL A TEST CASE FOR STRUCTURE-BASED FUNCTIONAL JRNL TITL 2 ASSIGNMENT: THE 1.2 A CRYSTAL STRUCTURE OF THE JRNL TITL 3 YJGF GENE PRODUCT FROM ESCHERICHIA COLI JRNL REF PROTEIN SCI. V. 8 2428 1999 |
In this file we find a series of water molecules with occupancies that suggest that they are located at special positions. In the box below I have taken only the waters from 1QU9 that have an occupancy other than 1.0.
HETATM 2986 O HOH A 345 36.123 72.223 49.938 0.50 35.86 O HETATM 3154 O HOH B 545 72.223 49.938 36.123 0.50 35.86 O HETATM 3313 O HOH C 745 49.938 36.123 72.223 0.50 35.86 O HETATM 2902 O HOH A 256 36.121 69.677 72.242 0.50 18.29 O HETATM 3068 O HOH B 456 69.677 72.242 36.121 0.50 18.29 O HETATM 3229 O HOH C 656 72.242 36.121 69.677 0.50 18.29 O HETATM 2928 O HOH A 284 39.241 72.228 72.228 0.50 23.81 O HETATM 3095 O HOH B 484 72.228 72.228 39.241 0.50 23.81 O HETATM 3255 O HOH C 684 72.228 39.241 72.228 0.50 23.81 O HETATM 2950 O HOH A 306 47.693 80.499 51.736 0.50 27.31 O HETATM 3117 O HOH B 506 80.499 51.736 47.693 0.50 27.31 O HETATM 3277 O HOH C 706 51.736 47.693 80.499 0.50 27.31 O HETATM 3014 O HOH B 330 43.028 43.028 43.028 0.33 32.19 O HETATM 3139 O HOH B 530 43.028 43.028 43.028 0.33 32.19 O HETATM 3169 O HOH B 730 43.028 43.028 43.028 0.33 32.19 O HETATM 3007 O HOH B 206 45.290 45.290 45.290 0.33 8.18 O HETATM 2998 O HOH A 406 45.290 45.290 45.290 0.33 8.18 O HETATM 3166 O HOH B 606 45.290 45.290 45.290 0.33 8.18 O HETATM 3010 O HOH B 276 54.593 54.593 54.593 0.33 21.40 O HETATM 3088 O HOH B 476 54.593 54.593 54.593 0.33 21.40 O HETATM 3168 O HOH B 676 54.593 54.593 54.593 0.33 21.40 O |
The CRYST and SCALE cards indeed indicate the existence of 2-fold and 3-fold axes.
CRYST1 72.220 72.220 72.220 90.00 90.00 90.00 P 2 2 2 12 SCALE1 0.013847 0.000000 0.000000 0.00000 SCALE2 0.000000 0.013847 0.000000 0.00000 SCALE3 0.000000 0.000000 0.013847 0.00000 |
So I can somewhat understand some of the the waters with occupancy 0.5. Some other waters with occupancy 0.5, however, seem at a first glance to not be exactly on a 2-fold axis. The atoms with occupancy 0.33 indeed seem positioned exactly at a 3-fold axis as the view below down that 3-fold axis seems to suggest. If you put a water with occupancy 0.33 on a 3-fold axis, there should not be three instances of that water, not even with different chain identifiers.
I am not such a hero with spacegroups and spacegroup symmetry, but I am a bit suprised this spacegroup is called P222 and not P23, but that is a different story.
A second note is that the spacegroup once was P23, but that the PDB expanded the file and made if P222, so it doesn't seem overly unlikely that the PDB is to blame for this problem.
The resolution of 1QU9, by the way, is 1.2 Ångström, so there should be enough data to do things well.

|
Waters with non-1.0 occupancy shown in a wireframe cell of 1QU9. |
EU name: 1VR1
(Date: Aug 24 2016 1VR1 )
Occupancies other than 1.0 are used when atoms are found at different locations either at different times or in different asymmetric units. Often though, they are also used to indicate that atoms are missing, invisible, or otherwise doubtful.
JRNL AUTH R.J.DEKKER,A.EICHINGER,A.A.STOOP,W.BODE, JRNL AUTH 2 H.PANNEKOEK,A.J.G.HORREVOETS JRNL TITL THE VARIABLE REGION-1 FROM TISSUE-TYPE PLASMINOGEN JRNL TITL 2 ACTIVATOR CONFERS SPECIFICITY FOR PLASMINOGEN JRNL TITL 3 ACTIVATOR INHIBITOR-1 TO THROMBIN BY FACILITATING JRNL TITL 4 CATALYSIS: RELEASE OF A KINETIC BLOCK BY A JRNL TITL 5 HETEROLOGOUS PROTEIN SURFACE LOOP JRNL REF J.MOL.BIOL. V. 293 613 1999 |
In the file 1VR1, though, we find some funny zero occupancies:
ATOM 996 N GLU H 97A 28.200 -13.994 26.148 1.00 42.09 N ATOM 997 CA GLU H 97A 27.495 -14.593 27.279 1.00 42.24 C ATOM 998 C GLU H 97A 26.054 -14.119 27.395 1.00 47.57 C ATOM 999 O GLU H 97A 25.064 -14.814 27.268 1.00 49.99 O ATOM 1000 CB GLU H 97A 28.231 -14.164 28.561 1.00 44.76 C ATOM 1001 CG GLU H 97A 28.577 -13.500 28.894 0.00 37.87 C ATOM 1002 CD GLU H 97A 29.575 -12.878 30.251 1.00 58.21 C ATOM 1003 OE1 GLU H 97A 28.669 -13.209 31.495 0.00 38.48 O ATOM 1004 OE2 GLU H 97A 30.490 -13.001 30.300 0.00 38.47 O HETATM 2396 O HOH H 313 28.850 -13.227 29.373 1.00 81.71 O |
I added one water to this box. One doesn't need a calculator to see that this water resides 'inside' the glutamate. The distance of this water to the Cγ and Cδ, respectively is 0.6 Ångström and 1.2 Ångström.
|
|