Ligands
WHAT_CHECK needs to 'understand' all residues and ligands in terms of topology and
hydrogen bond donor/acceptor behaviour. WHAT_CHECK tries to interpret non-canonical
(i.e. not one of the 20 normal ones) amino acids itself, and does a rather poor job on that,
still. So, if non-canonical amino acids are detected, then the validation of their direct
environment, especially when hydrogen bonds are involved, should be taken with
a grain of salt. WHAT IF uses the Dundee PRODRUG software for the interpretation of
ligands/drugs/ lipids, etc. If such a small molecule is encountered, and PRODRUG cannot
produce a WHAT_CHECK topology for it, then WHAT_CHECK will have difficulties with the
validation of that small molecule and its direct environment, especially when hydrogen
bonds are involved.
Prodrug
Prodrug was started by Daan van Aalten when he did an internship (many many years ago) in our
group at the EMBL. He later continued working on the software when he got his own group in
Dundee. More recently, Alexander Schuetelkopf in Daan's group took over the maintenance.
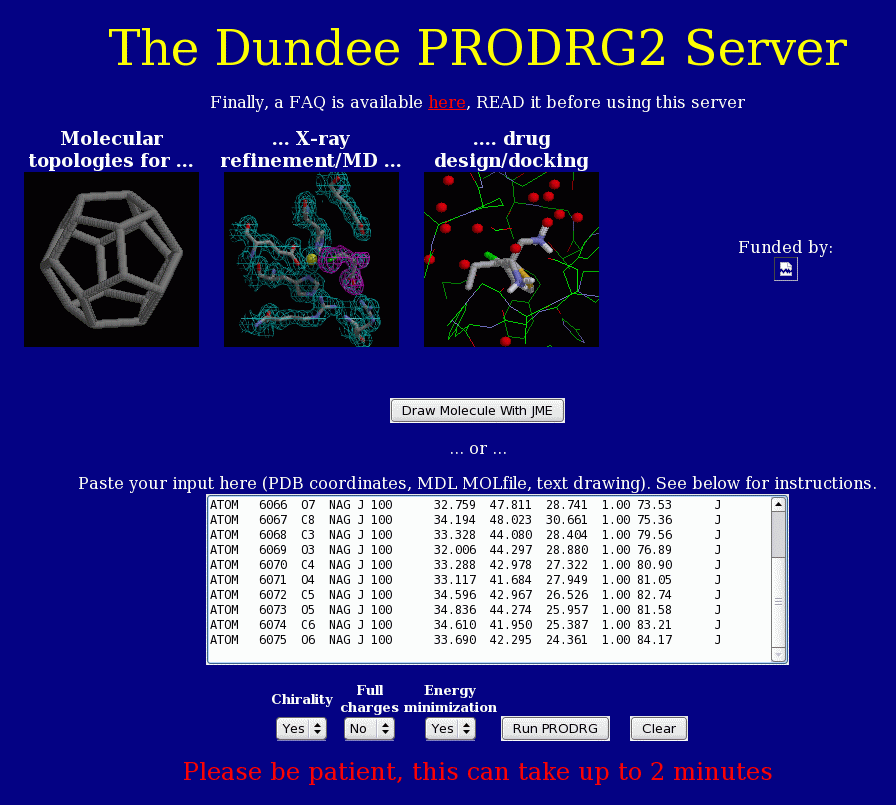
|
Figure 3. Prodrug's interactive interface. WHAT IF has Prodrug built-in so as a user you
don't have to worry about these topology problems, unless there is a problem with the
topology generation, of course. The note below shows one topology generation example for
the curious ones among you. (Prodrug home).
|
Supplemental material
Normally, you don't have to worry about these things, but if you are solving
a structure with a ligand/drug in it, and things seem to be going wrong
somehow, then you can use Prodrug to check the topology entry for that
ligand/drug in your refinement software.
