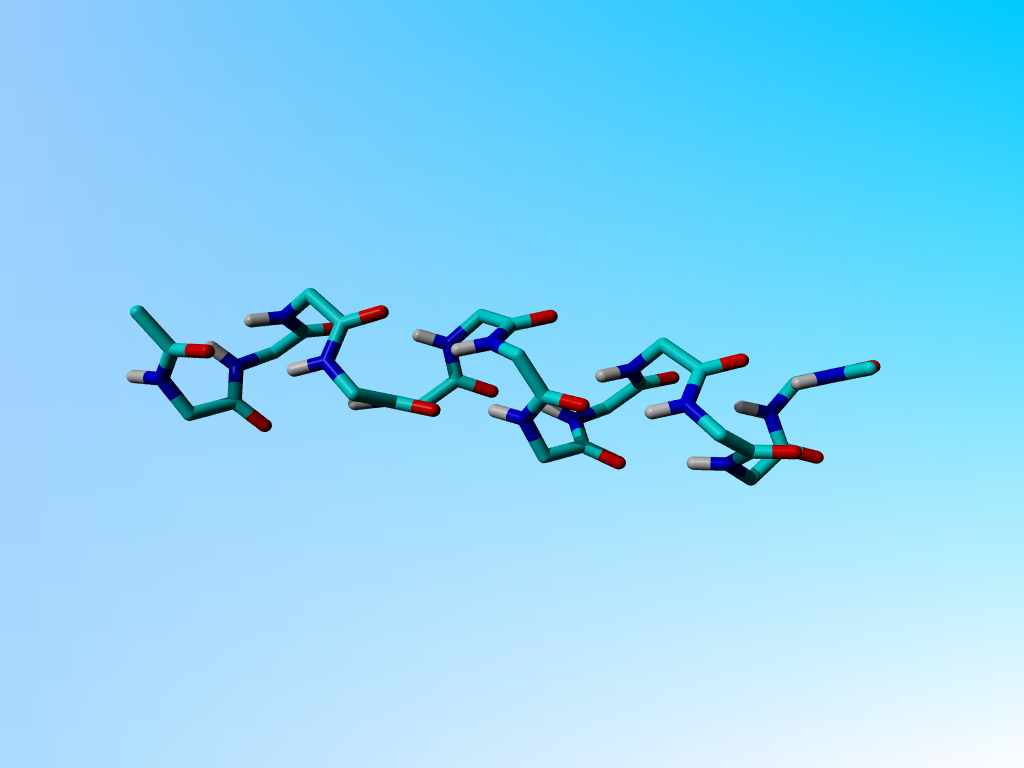
The 1cek poly-gly helix. Hydrophobic hydrogens were added in the movie that you get when you click on the picture because Yasare refuses to calculate hydrogen bonds without them present.
There is no doubt that, in general, it is a very good idea to deposite structures and the underlying experimental data. This was made very clear in:
PDB Improvement Starts with Data Deposition Joosten and Vriend Science 13 July 2007: 195-196 DOI: 10.1126/science.317.5835.195 |
and in dozens of other articles. However, there are exceptions; like fraud. We will come back to fraud at some later stage. However, there are also structures that contain no information.
JRNL AUTH S.J.OPELLA,F.M.MARASSI,J.J.GESELL,A.P.VALENTE, JRNL AUTH 2 Y.KIM,M.OBLATT-MONTAL,M.MONTAL JRNL TITL STRUCTURES OF THE M2 CHANNEL-LINING SEGMENTS FROM JRNL TITL 2 NICOTINIC ACETYLCHOLINE AND NMDA RECEPTORS BY NMR JRNL TITL 3 SPECTROSCOPY JRNL REF NAT.STRUCT.BIOL. V. 6 374 1999 |
1CEK is an example of a structure that contains no information. We are not saying anything bad about Stan Opella and his people who solved this structure because solving protein structures with solid state NMR was in 1999 a big thing, and proving that this peptide is helical also added something to the world's body of knowledge.
However, the authors couldn't see any side chains in the structure, and the data did not allow them to get any precision. If you look at the structure, you see a badly distorted poly-glycine helix:

|
The 1cek poly-gly helix. Hydrophobic hydrogens were added in the movie that you get when you click on the picture because Yasare refuses to calculate hydrogen bonds without them present. |
The authors write themselves:
REMARK 7 THE STRUCTURE MUST BE VIEWD AS A SUPRAMOLECULAR ASSEMBLY REMARK 7 TOGETHER WITH THE LIPID BILAYER MEMBRANE. SOLID-STATE NMR REMARK 7 OF PROTEINS IN ORIENTED BILAYERS GIVES STRUCTURES WITH REMARK 7 COORDINATES THAT ARE FIXED RELATIVE TO THE BILAYER AND REMARK 7 CONTAIN BOTH STRUCTURE AND GLOBAL TOPOLOGY INFORMATION. REMARK 7 ACHR M2 IS A TRANSMEMBRANE, AMPHIPATHIC ALPHA-HELIX. THE REMARK 7 LONG HELIX AXIS IS TILTED 12 DEGRESS FROM THE BILAYER REMARK 7 NORMAL. THE HELIX IS ROTATED AROUND ITS LONG AXIS SUCH THAT REMARK 7 THE POLAR RESIDUES FACE THE N-TERMINAL SIDE OF THE REMARK 7 MEMBRANE. |
I have no idea what is the N-terminal side of the membrane... Anyway, this text and the secondary structure determination:
1 - 14 AISVLLAQAVFLLL
1 - 14 ? HHHHHHHHHH ?
|
were enough to fully describe this structure.
EU name: 2PDE
(Date: Aug 24 2016 2PDE )
In 1993 it probably wasn't easy yet to solve a structure by NMR. But molecular visualisation software did exist! WHAT IF, for example, was already 6 years old at that time. I suggest 2PDE is removed from the PDB so that the article about its high-resolution structure can be removed from our memories.
JRNL AUTH Y.N.KALIA,S.M.BROCKLEHURST,D.S.HIPPS,E.APPELLA, 2PDE 14 JRNL AUTH 2 K.SAKAGUCHI,R.N.PERHAM 2PDE 15 JRNL TITL THE HIGH RESOLUTION STRUCTURE OF THE PERIPHERAL 2PDE 16 JRNL TITL 2 SUBUNIT-BINDING DOMAIN OF DIHYDROLIPOAMIDE 2PDE 17 JRNL TITL 3 ACETYLTRANSFERASE FROM THE PYRUVATE DEHYDROGENASE 2PDE 18 JRNL TITL 4 MULTIENZYME COMPLEX OF BACILLUS 2PDE 19 JRNL TITL 5 STEAROTHERMOPHILUS 2PDE 20 JRNL REF J.MOL.BIOL. V. 230 323 1993 2PDE 21 JRNL REFN ASTM JMOBAK UK ISSN 0022-2836 0070 2PDE 22 |
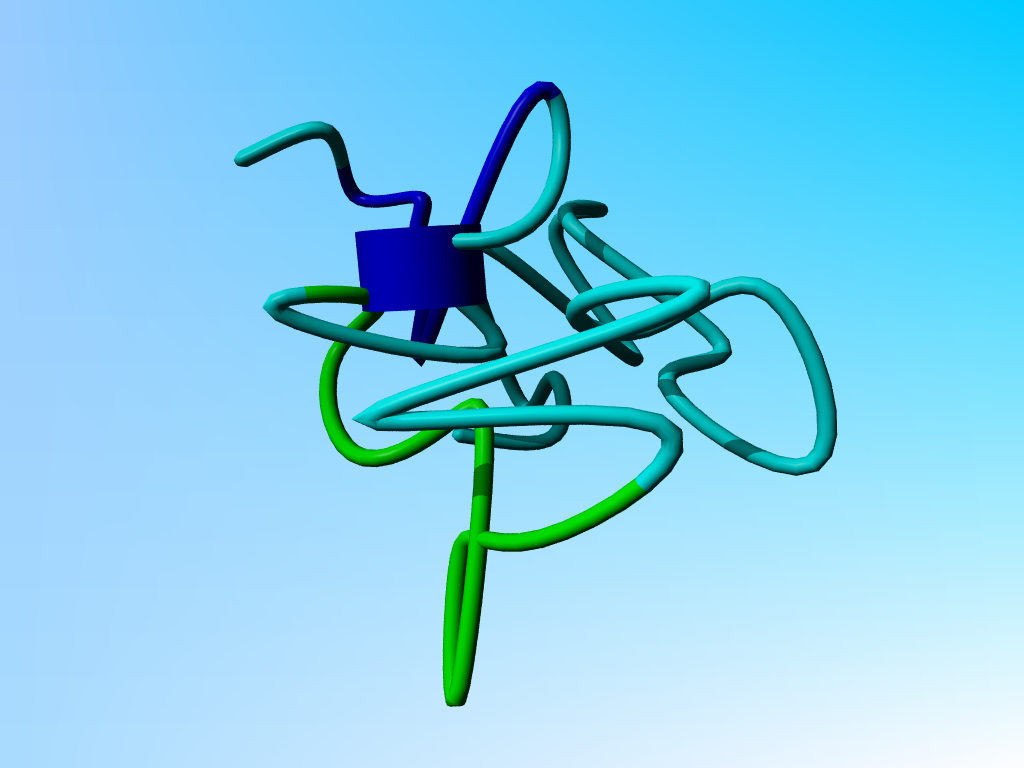
The PDB file 2PDE is best described by a short movie:
|
The funny blue cylinder (normally used to indicate a helix) here represents a chaotic attractor. The movie is 20M big, but the laugh is worth the wait. |