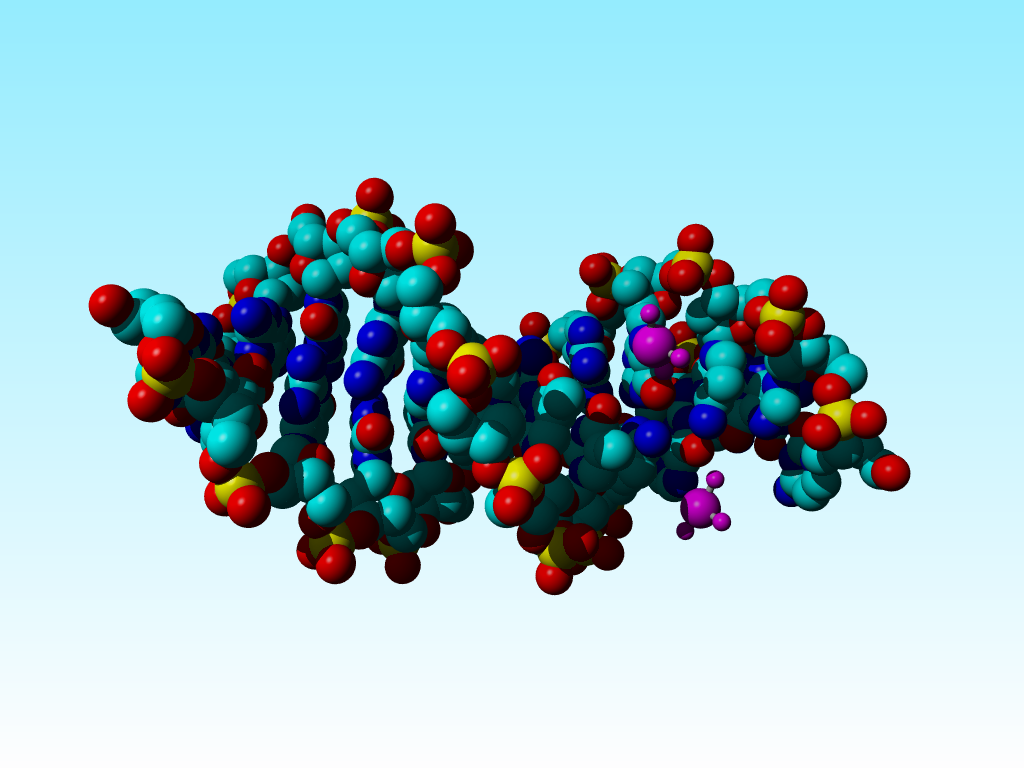
The PDB file 5bna. Waters were removed. The cisplatins are in purple. This picture is also available rotating around the horizontal axis.
Things have names so you can refer to them, search for them in a database, etcetera. If your kids would be called John and Mary when at home, but Harry and Jane at school, and Martin and Jo when on vacation, you would be hard pressed as a parent to at all times call them by the right name.
Such simple logic does not hold for the PDB. In the PDB it is even possible to change names of items from one line to the next. Look at:
HETNAM GLB BETA-D-GALACTOSE HETNAM GLA ALPHA D-GALACTOSE |
which I found in the file 1OKO. Why is there a - between BETA and D, but not
between ALPHA and D? This looks a stupid little thing, but it adds an extra
layer of complication for the software engineer who wants to generate a search
engine that can find ligands in PDB files.
EU name: 5BNA
(Date: Aug 24 2016 5BNA )
JRNL AUTH R.M.WING,P.PJURA,H.R.DREW,R.E.DICKERSON 5BNAB 2 JRNL TITL THE PRIMARY MODE OF BINDING OF CISPLATIN TO A 5BNAB 3 JRNL TITL 2 B-/DNA$ DODECAMER. 5BNAB 4 JRNL TITL 3 C-*G-*C-*G-*A-*A-*T-*T-*C-*G-*C-*G 5BNAB 5 JRNL REF /EMBO$ J. V. 3 1201 1984 5BNAB 6 |
Often, problems occur for which a solution is hard to imagine. Take 5bna. This structure holds cisplatin. Unfortunately, the individual atoms attached to the platinum could not be distinguished:
REMARK 5 THE TWO STRANDS IN THE ASYMMETRIC UNIT HAVE BEEN ASSIGNED 5BNA 59 REMARK 5 CHAIN IDENTIFIERS *A* AND *B*. IN THIS ENTRY THE RESIDUES 5BNA 60 REMARK 5 IN EACH CHAIN ARE NUMBERED FROM 1 TO 12. IN THE DATA AS 5BNA 61 REMARK 5 ORIGINALLY DEPOSITED THE RESIDUES IN THE *B* CHAIN WERE 5BNA 62 REMARK 5 NUMBERED FROM 13 TO 24. THE COORDINATES FOR THREE 5BNA 63 REMARK 5 CISPLATIN GROUPS ARE LISTED FOLLOWING THE B-CHAIN. THE 5BNA 64 REMARK 5 CISPLATIN MOLECULES ARE ASSUMED TO BE IN A MODIFIED FORM IN 5BNA 65 REMARK 5 WHICH THE TWO CHLORINE ATOMS HAVE BEEN REPLACED, ONE BY A 5BNA 66 REMARK 5 WATER MOLECULE AND THE OTHER BY THE N7 ATOM OF A GUANINE TO 5BNA 67 REMARK 5 WHICH IT IS COVALENTLY BONDED. THE WATER AND THE TWO 5BNA 68 REMARK 5 NITROGEN ATOMS BONDED TO THE PLATINUM HAVE NOT BEEN 5BNA 69 REMARK 5 DISTINGUISHED AND ARE IDENTIFIED AS A1, A2, AND A3. CPT 25 5BNA 70 REMARK 5 IS BONDED TO G B 4, CPT 26 TO G A 4, AND CPT 27 TO G A 10. 5BNA 71 |
So the atoms got atomtype A:
HETATM 497 PT CPT 25 15.894 14.427 2.341 1.00 29.11 1 5BNA 594 HETATM 498 A1 CPT 25 17.492 13.427 3.062 1.00 0.00 2 5BNA 595 HETATM 499 A2 CPT 25 14.637 15.234 1.311 1.00 19.23 5BNA 596 HETATM 500 A3 CPT 25 14.805 12.857 1.752 1.00 0.00 2 5BNA 597 HETATM 501 PT CPT 26 20.276 22.935 14.540 1.00 31.23 1 5BNA 598 HETATM 502 A1 CPT 26 21.408 21.700 14.028 1.00 0.00 2 5BNA 599 HETATM 503 A2 CPT 26 18.763 24.527 15.013 1.00 36.54 5BNA 600 HETATM 504 A3 CPT 26 21.733 24.620 14.431 1.00 0.00 2 5BNA 601 HETATM 505 PT CPT 27 6.884 19.581 -3.252 1.00 20.79 1 5BNA 602 HETATM 506 A1 CPT 27 5.598 20.789 -2.761 1.00 0.00 2 5BNA 603 HETATM 507 A2 CPT 27 8.523 18.347 -3.721 1.00312.44 5BNA 604 HETATM 508 A3 CPT 27 5.863 17.708 -3.342 1.00 0.00 2 5BNA 605 |
And although this seems better that randomly guessing which of the atoms is what, most molecular visualizer don't like the Mendeleev symbol A very much.

|
The PDB file 5bna. Waters were removed. The cisplatins are in purple. This picture is also available rotating around the horizontal axis. |
In the beginning of the previous century, when the format of the PDB was defined, the constraints on the format were partly set by punch-cards, and partly by the limited imagination of the people at that moment. Consequently, they reserved only 4 characters for the atom names. And every 5-10 years this causes a major PDB-overhaul/catastrophe. We need 5 characters and in the future perhaps 6 to properly define atom names.
At the latest overhaul, in 2006-2007 it was decided to do away with the rule that the first two positions of the 4 characters reserved for the atom name should be the Mendeleev symbol. Instead, all 4 characters could be used at will, and the columns 77-78 were to be used for the Mendeleev symbol.
The PDB undoubtedly used a big and complicated script to make sure that those Mendeleev symbols were placed there properly. But in 2AXT this script seems to have failed. I can imagine finding Neodymium in proteins, but Neon baffles me a bit... Undoubtedly, they forgot to include UNK in the script.
1 2 3 4 5 6 7
123456789012345678901234567890123456789012345678901234567890123456789012345678
ATOM 19239 N UNK X 3 23.324 78.683 82.548 1.00 94.35 N
ATOM 19240 CA UNK X 3 22.200 79.572 82.218 1.00 93.04 C
ATOM 19241 C UNK X 3 22.207 79.973 80.744 1.00 92.07 C
ATOM 19242 O UNK X 3 23.202 79.740 80.052 1.00 92.45 O
ATOM 19243 CB UNK X 3 22.248 80.840 83.088 1.00 92.62 C
ATOM 19244 CG UNK X 3 20.979 81.683 82.968 1.00 91.97 C
ATOM 19245 OD1 UNK X 3 19.935 81.341 83.545 1.00 90.79 O
ATOM 19246 ND2 UNK X 3 21.059 82.776 82.200 1.00 91.80 ND
ATOM 19247 N UNK X 4 21.100 80.558 80.266 1.00 90.70 N
...
ATOM 19264 CB UNK X 6 25.712 79.333 77.369 1.00 78.84 C
ATOM 19265 CG UNK X 6 26.291 78.771 78.645 1.00 78.32 C
ATOM 19266 CD UNK X 6 25.709 77.423 79.000 1.00 78.34 C
ATOM 19267 OE1 UNK X 6 24.535 77.162 78.731 1.00 77.93 O
ATOM 19268 NE2 UNK X 6 26.511 76.565 79.630 1.00 76.34 NE
|
EU name: URANYL
(Date: Aug 24 2016 URANYL )
JRNL AUTH U.HOFFMULLER,T.KNAUTE,M.HAHN,W.HOHNE, JRNL AUTH 2 J.SCHNEIDER-MERGENER,A.KRAMER JRNL TITL EVOLUTIONARY TRANSITION PATHWAYS FOR CHANGING JRNL TITL 2 PEPTIDE LIGAND SPECIFICITY AND STRUCTURE. JRNL REF EMBO J. V. 19 4866 2000 |
Sometimes the crystallographer cannot see the C-terminal N residues. Nothing wrong with that. The PDB, at some time, and somewhat surprisingly decided to let the last residue have a C-terminal extra N, instead of nothig or an O, to indicate that the last residue in the PDB file in reality is not the last residue in the file. Except for the stealthy way of introduction, this wasn't a very bad concept.
And then cam 1HH6. In this PDB file the C-chain ends with the famous atom U-nk which, as the Mendeleev symbol really is U, must be read as uranyl-nk.
.... ATOM 3348 N LEU C 11 2.409 43.575 143.138 1.00 84.40 N ATOM 3349 CA LEU C 11 2.357 44.193 141.811 1.00 85.61 C ATOM 3350 C LEU C 11 2.408 45.720 141.867 1.00 85.79 C ATOM 3351 O LEU C 11 1.436 46.370 142.243 1.00 45.92 O ATOM 3352 CB LEU C 11 1.091 43.745 141.062 1.00 86.19 C ATOM 3353 CG LEU C 11 1.046 42.395 140.318 1.00 87.35 C ATOM 3354 CD1 LEU C 11 1.351 41.233 141.256 1.00 87.35 C ATOM 3355 CD2 LEU C 11 -0.336 42.217 139.689 1.00 87.35 C ATOM 3356 UNK LEU C 11 3.550 46.289 141.495 1.00 86.07 U TER 3357 LEU C 11 .... |
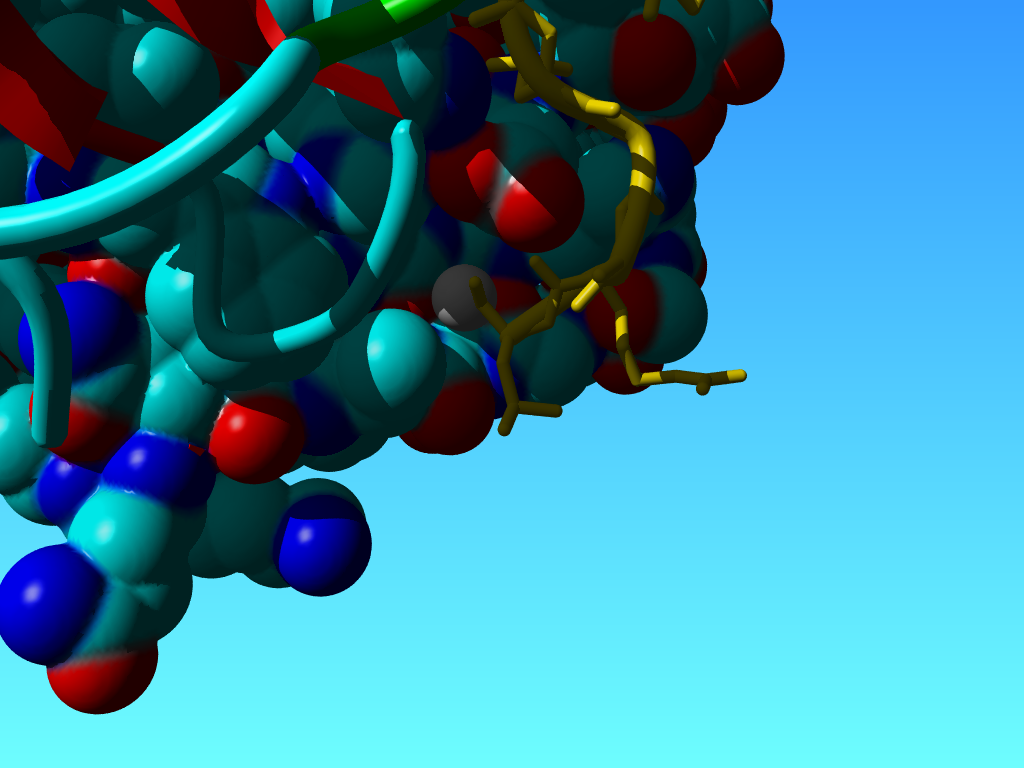
One can understand how such an administrative incovenience gets born, but when I look at the structure and see the U fully buried, I stop to understand:
|
The molecular surface shown on the B chain. The C-chain in yellow. The U at the C-terminal end of the C-chain is shown as a gray ball. |
EU name: ETA
(Date: Aug 24 2016 ETA )
Sometimes it looks as if the depositors don't want us to properly read and parse their files. Take 1AV2, for example.
JRNL AUTH B.M.BURKHART,N.LI,D.A.LANGS,W.A.PANGBORN,W.L.DUAX JRNL TITL THE CONDUCTING FORM OF GRAMICIDIN A IS A JRNL TITL 2 RIGHT-HANDED DOUBLE-STRANDED DOUBLE HELIX. JRNL REF PROC.NATL.ACAD.SCI.USA V. 95 12950 1998 |
1AV2 holds a lot of non-canonical amino acids like D-leucine and D-valine. It also holds ethanolamine, with as entity name (residue code) ETA:
HETATM 263 CA ETA A 16 27.320 17.142 24.093 1.00 26.18 C HETATM 264 N ETA A 16 26.386 18.262 24.001 1.00 21.37 N HETATM 265 CB ETA A 16 28.575 17.453 23.302 1.00 32.91 C HETATM 266 O ETA A 16 29.311 16.206 23.515 1.00 45.04 O |
This ETA also holds several protons. One must actually read the proton names to see that this is actually ethanolamine and not a glycine with the second C-terminal oxygen missing and the C misnamed. Why does ethanol need a CA and CB, why cannot it be C1 and C2? Beats me, and till yesterday my software.