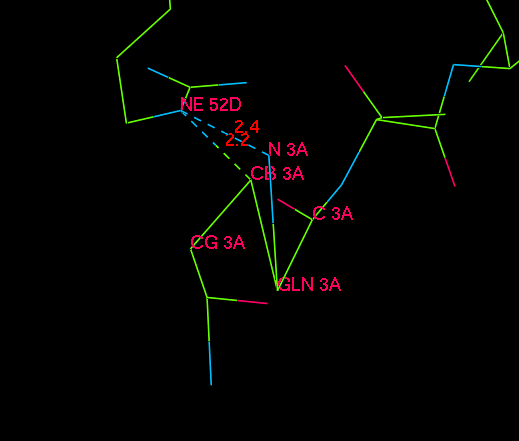
The direct environment of the rather funny residue 1. I indicated two very short distances to the Arginine.
JRNL AUTH N.RIFFEL,K.HARLOS,O.IOURIN,Z.RAO,A.KINGSMAN, JRNL AUTH 2 D.STUART,E.FRY JRNL TITL ATOMIC RESOLUTION STRUCTURE OF MOLONEY MURINE JRNL TITL 2 LEUKAEMIA VIRUS MATRIX PROTEIN AND ITS JRNL TITL 3 RELATIONSHIP TO OTHER RETROVIRAL MATRIX PROTEINS. JRNL REF STRUCTURE V. 10 1627 2002 JRNL REFN ASTM STRUE6 UK ISSN 0969-2126 |
In many proteins we find totally knotted or otherwise distorted N-terminal residues. Here is one example in 1mn8:

|
The direct environment of the rather funny residue 1. I indicated two very short distances to the Arginine. |
Clear, the B-factors of this residue are a bit high:
ATOM 1 N GLN A 3 1.487 34.339 -28.185 1.00 56.65 N ATOM 2 CA GLN A 3 0.352 32.071 -28.036 1.00 52.23 C ATOM 3 C GLN A 3 1.691 33.044 -27.146 1.00 53.94 C ATOM 4 O GLN A 3 1.212 33.971 -26.510 1.00 67.04 O ATOM 5 CB GLN A 3 0.910 34.072 -28.502 1.00 58.09 C ATOM 6 CG GLN A 3 -0.929 34.047 -27.176 1.00 71.02 C ATOM 7 CD GLN A 3 -1.088 33.260 -26.417 1.00 67.98 C ATOM 8 OE1 GLN A 3 -0.012 32.867 -24.786 1.00 65.10 O ATOM 9 NE2 GLN A 3 -1.856 31.906 -25.959 1.00 81.68 N |
but that should not be an excuse for a residue in which the bond-lengths are longer than the atomic distances to other residues. This structure was solved in 2002 at 1.0 Ångström resolution...